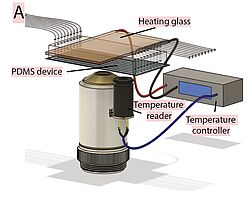
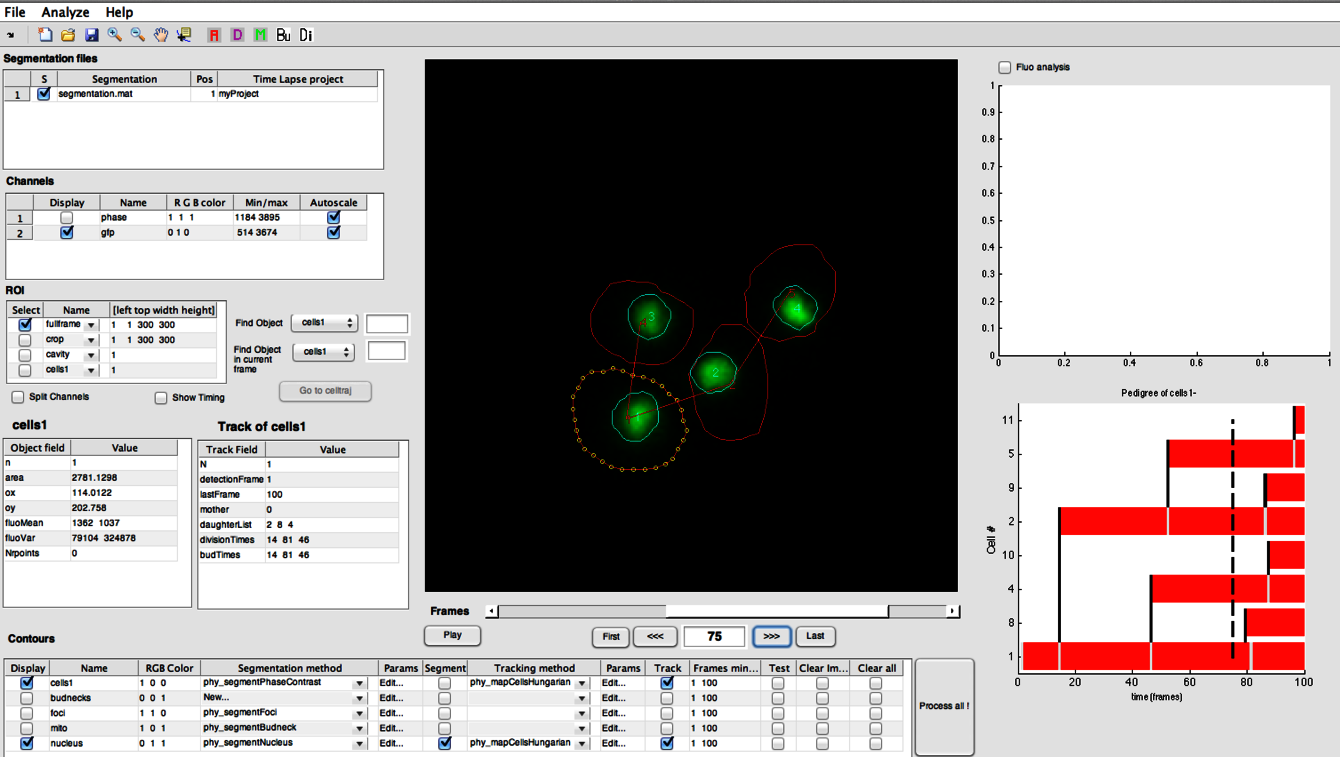
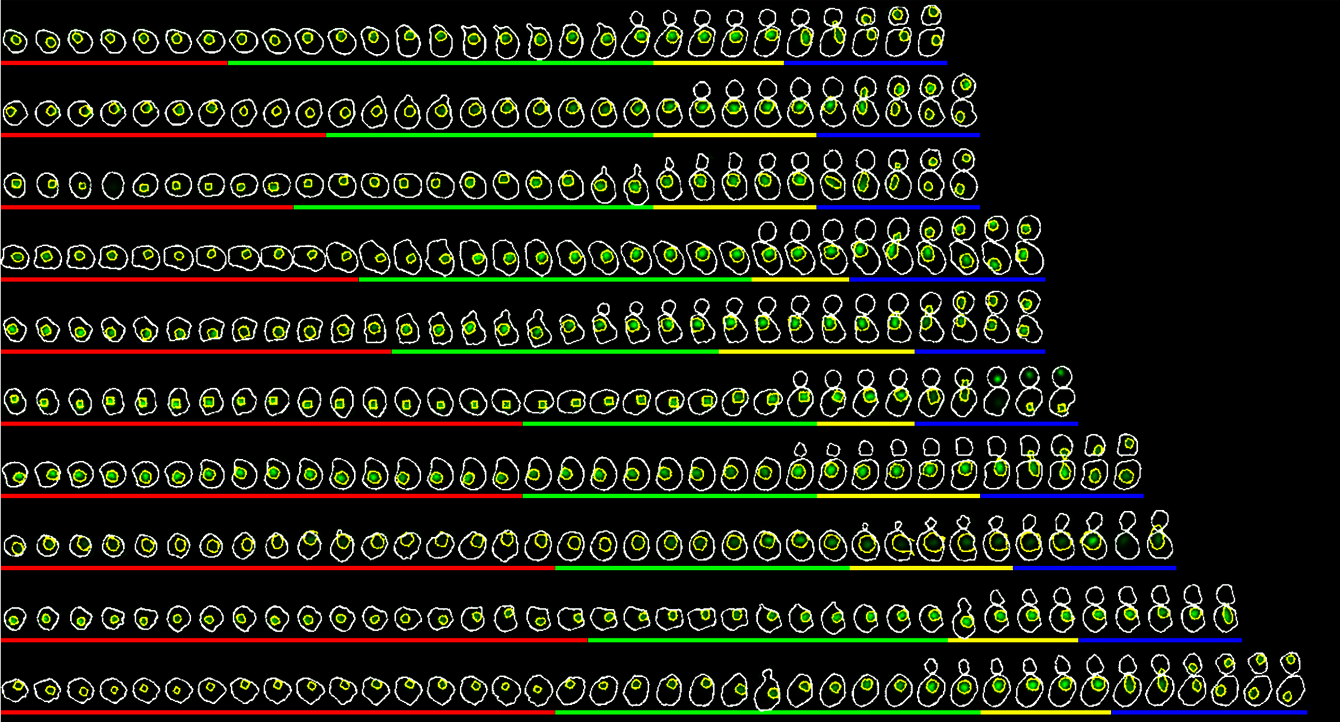
Laboratory equipment and techniques
The team uses classical methods of yeast molecular genetics (transformation, crosses, etc.) to generate mutant strains (auxin-dependent degron, conditional mutants, conditional expression) and fluorescent markers (redox or pH ratiometric probes, fusion proteins, fluorescent timers, etc.).